This web page was produced as an assignment for Gen677 at UW-Madison Spring 2010
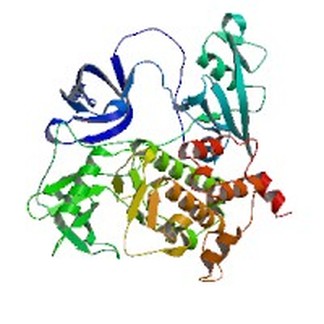
ABL1 protein
Full protein name: Tyrosine-protein kinase ABL1
Amino acid length: 1130 aa
Protein accession number: P00519
Isoelectric point: 8.84
Molecular weight: 122872.56
| abl1_protein_sequence.rtf |
Function
The ABL1 protein plays a large role in regulating processes linked to cell growth and survival throughout majority of the tissues in the body. Some of these processes include cytoskeleton remodeling during cell differentiation, cell division, and cell adhesion and regulation of DNA repair through the activation of the proapoptotic pathway. Majority of these regulations occur through protein-protein interactions.
Isoforms
There are two known isoforms of the ABL protein: Isoform A and Isoform B. The difference between the two isoforms lies within the beginning of the amino acid sequence. Using Isoform A as the canonical sequence, the two isoforms differ in the first 25 amino acid sequence. Isoform B also has an additional 19 amino acids inserted between the 12 and 13 position. Alternative splicing is the reason for the differences between the two isoforms.
Post-transcriptional Modifications
The main post-transcriptional modifications as indicated by UniProt was phosphorylation by PRKDC and ATM. The phosphorylation occurs on Ser-446 for DNA damage-induced activation and on Thr-735 for cytoplasmic translocation. (4) Isoform B is also myristoylated on Gly-2. (4) No post-transcriptional modifications could be found using Expasy.
Subcellular Location
The ABL1 protein is located in the cytoplasm of the cell, specifically the cytoskeleton, and the nucleus. Isoform B is also located in association with the nuclear membrane as a lipid anchor.
For more details about the ABL1 protein, click on the links below.
Homologs
Domains
Protein Interactions
References
1. http://www.ncbi.nlm.nih.gov
2. ModBase: Database of Comparative Protein Structure Models http://modbase.compbio.ucsf.edu/modbase-cgi/index.cgi
3. Expasy http://us.expasy.org/tools/pi_tool.html
4. UniProt http://www.uniprot.org/uniprot/P00519#P00519-2
2. ModBase: Database of Comparative Protein Structure Models http://modbase.compbio.ucsf.edu/modbase-cgi/index.cgi
3. Expasy http://us.expasy.org/tools/pi_tool.html
4. UniProt http://www.uniprot.org/uniprot/P00519#P00519-2